Examples in Summary and
Analysis of Extension Program Evaluation
SAEPER: Goodness-of-Fit Tests for Nominal Variables
Packages used in this chapter
The following commands will install these packages
if they are not already installed:
if(!require(dplyr)){install.packages("dplyr")}
if(!require(ggplot2)){install.packages("ggplot2")}
if(!require(grid)){install.packages("grid")}
if(!require(pwr)){install.packages("pwr")}
See the Handbook for information on these topics.
How the test works
Chi-square goodness-of-fit example
###
--------------------------------------------------------------
### Drosophila example, Chi-square goodness-of-fit, p. 46
### --------------------------------------------------------------
observed = c(770, 230) # observed
frequencies
expected = c(0.75, 0.25) # expected
proportions
chisq.test(x = observed,
p = expected)
X-squared = 2.1333, df = 1, p-value = 0.1441
# # #
See the Handbook for information on these topics.
Examples: extrinsic hypothesis
### --------------------------------------------------------------
### Crossbill example, Chi-square goodness-of-fit, p. 47
### --------------------------------------------------------------
observed = c(1752, 1895) # observed frequencies
expected = c(0.5, 0.5) # expected proportions
chisq.test(x = observed,
p = expected)
X-squared = 5.6071, df = 1, p-value = 0.01789
# # #
###
--------------------------------------------------------------
### Rice example, Chi-square goodness-of-fit, p. 47
### --------------------------------------------------------------
observed = c(772, 1611, 737)
expected = c(0.25, 0.50, 0.25)
chisq.test(x = observed,
p = expected)
X-squared = 4.1199, df = 2, p-value = 0.1275
# # #
###
--------------------------------------------------------------
### Bird foraging example, Chi-square goodness-of-fit, pp. 47–48
### --------------------------------------------------------------
observed = c(70, 79, 3, 4)
expected = c(0.54, 0.40, 0.05, 0.01)
chisq.test(x = observed,
p = expected)
X-squared = 13.5934, df = 3, p-value = 0.0035
# # #
Example: intrinsic hypothesis
###
--------------------------------------------------------------
### Intrinsic example, Chi-square goodness-of-fit, p. 48
### --------------------------------------------------------------
observed = c(1203, 2919, 1678)
expected.prop = c(0.211, 0.497, 0.293)
expected.count = sum(observed)*expected.prop
chi2 = sum((observed- expected.count)^2/ expected.count)
chi2
[1] 1.082646
pchisq(chi2,
df=1,
lower.tail=FALSE)
[1] 0.2981064
# # #
Graphing the results
The first example below will use the barplot function in the native graphics package to produce a simple plot. First we will calculate the observed proportions and then copy those results into a matrix format for plotting. We’ll call this matrix Matriz. See the “Chi-square Test of Independence” section for a few notes on creating matrices.
The second example uses the package ggplot2, and uses a data frame instead of a matrix. The data frame is named Forage. For this example, the code calculates confidence intervals and adds them to the data frame. This code could be skipped if those values were determined manually and put into a data frame from which the plot could be generated.
Sometimes factors will need to have the order of their levels specified for ggplot2 to put them in the correct order on the plot, as in the second example. Otherwise R will alphabetize levels.
Simple bar plot with barplot
###
--------------------------------------------------------------
### Simple bar plot of proportions, p. 49
### Uses data in a matrix format
### --------------------------------------------------------------
observed = c(70, 79, 3, 4)
expected = c(0.54, 0.40, 0.05, 0.01)
total = sum(observed)
observed.prop = observed / total
observed.prop
[1] 0.44871795 0.50641026 0.01923077 0.02564103
### Re-enter data as a matrix
Input =("
Value Douglas.fir Ponderosa.pine Grand.fir Western.larch
Observed 0.4487179 0.5064103 0.01923077 0.02564103
Expected 0.5400000 0.4000000 0.05000000 0.01000000
")
Matriz = as.matrix(read.table(textConnection(Input),
header=TRUE,
row.names=1))
Matriz
Douglas fir Ponderosa pine Grand fir Western larch
Observed 0.4487179 0.5064103 0.01923077 0.02564103
Expected 0.5400000 0.4000000 0.05000000 0.01000000
barplot(Matriz,
beside=TRUE,
legend=TRUE,
ylim=c(0, 0.6),
xlab="Tree species",
ylab="Foraging proportion")
# # #
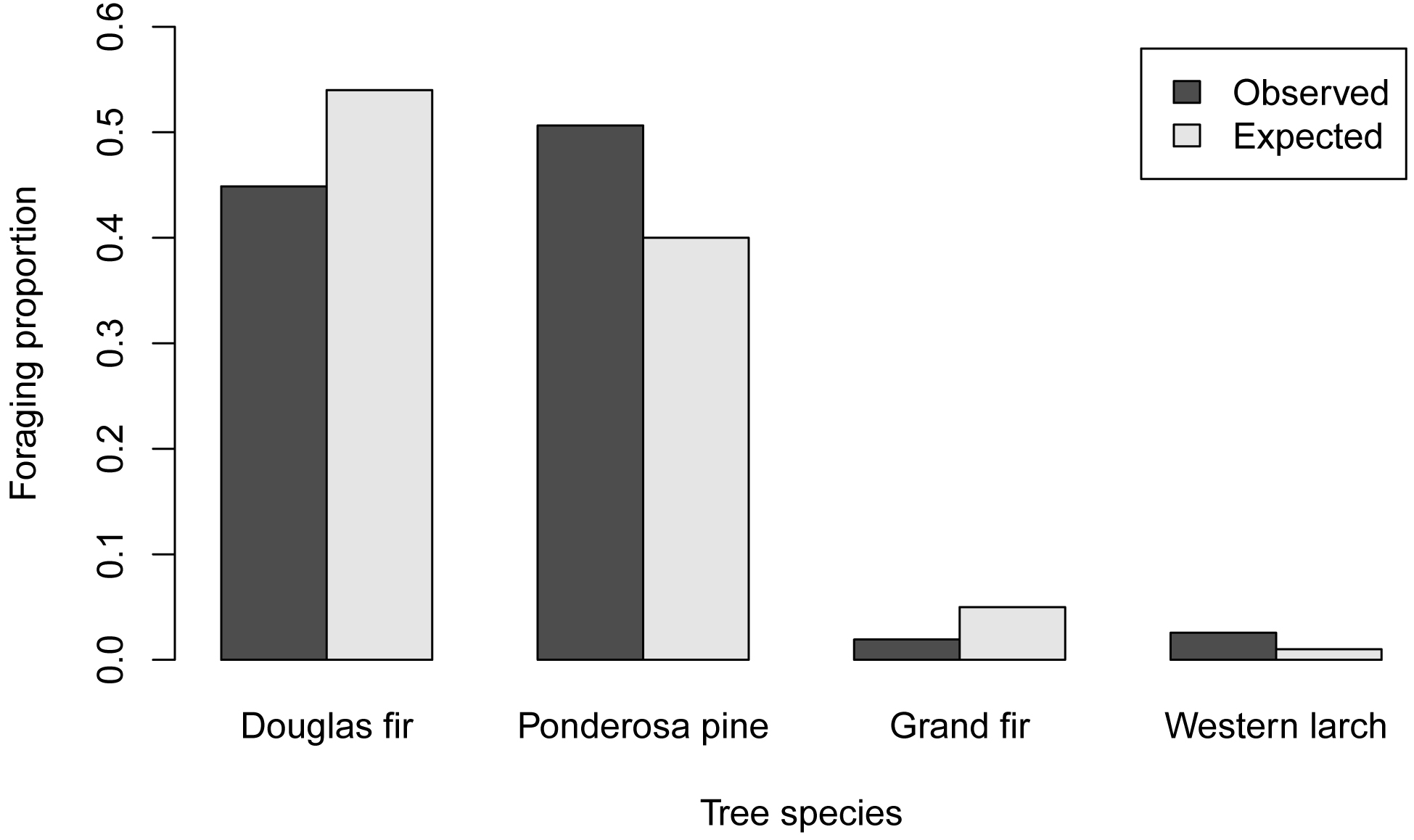
Bar plot with confidence intervals with ggplot2
The plot below is a bar char with confidence intervals. The code calculates confidence intervals. This code could be skipped if those values were determined manually and put in to a data frame from which the plot could be generated.
Sometimes factors will need to have the order of their levels specified for ggplot2 to put them in the correct order on the plot. Otherwise R will alphabetize levels.
### --------------------------------------------------------------
### Graph example, Chi-square goodness-of-fit, p. 49
### Using ggplot2
### Plot adapted from:
### shinyapps.stat.ubc.ca/r-graph-catalog/
### --------------------------------------------------------------
Input =("
Tree Value Count Total Proportion Expected
'Douglas fir' Observed 70 156 0.4487 0.54
'Douglas fir' Expected 54 100 0.54 0.54
'Ponderosa pine' Observed 79 156 0.5064 0.40
'Ponderosa pine' Expected 40 100 0.40 0.40
'Grand fir' Observed 3 156 0.0192 0.05
'Grand fir' Expected 5 100 0.05 0.05
'Western larch' Observed 4 156 0.0256 0.01
'Western larch' Expected 1 100 0.01 0.01
")
Forage = read.table(textConnection(Input),header=TRUE)
### Specify the order of factor levels. Otherwise R
will alphabetize them.
library(dplyr)
Forage =
mutate(Forage,
Tree = factor(Tree, levels=unique(Tree)),
Value = factor(Value, levels=unique(Value))
)
### Add confidence intervals
Forage =
mutate(Forage,
low.ci = apply(Forage[c("Count", "Total",
"Expected")],
1,
function(x)
binom.test(x["Count"], x["Total"],
x["Expected"]
)$ conf.int[1]),
upper.ci = apply(Forage[c("Count", "Total",
"Expected")],
1,
function(x)
binom.test(x["Count"], x["Total"],
x["Expected"]
)$ conf.int[2])
)
Forage$ low.ci [Forage$ Value == "Expected"] = 0
Forage$ upper.ci [Forage$ Value == "Expected"] = 0
Forage
Tree Value Count Total Proportion Expected low.ci upper.ci
1 Douglas fir Observed 70 156 0.4487 0.54 0.369115906 0.53030534
2 Douglas fir Expected 54 100 0.5400 0.54 0.000000000 0.00000000
3 Ponderosa pine Observed 79 156 0.5064 0.40 0.425290653 0.58728175
4 Ponderosa pine Expected 40 100 0.4000 0.40 0.000000000 0.00000000
5 Grand fir Observed 3 156 0.0192 0.05 0.003983542 0.05516994
6 Grand fir Expected 5 100 0.0500 0.05 0.000000000 0.00000000
7 Western larch Observed 4 156 0.0256 0.01 0.007029546 0.06434776
8 Western larch Expected 1 100 0.0100 0.01 0.000000000 0.00000000
### Plot adapted from:
### shinyapps.stat.ubc.ca/r-graph-catalog/
library(ggplot2)
library(grid)
ggplot(Forage,
aes(x = Tree, y = Proportion, fill = Value,
ymax=upper.ci, ymin=low.ci)) +
geom_bar(stat="identity", position = "dodge", width
= 0.7) +
geom_bar(stat="identity", position = "dodge",
colour = "black", width = 0.7,
show_guide = FALSE) +
scale_y_continuous(breaks = seq(0, 0.60, 0.1),
limits = c(0, 0.60),
expand = c(0, 0)) +
scale_fill_manual(name = "Count type" ,
values = c('grey80', 'grey30'),
labels = c("Observed value",
"Expected value")) +
geom_errorbar(position=position_dodge(width=0.7),
width=0.0, size=0.5, color="black") +
labs(x = "Tree species",
y = "Foraging proportion") +
## ggtitle("Main title") +
theme_bw() +
theme(panel.grid.major.x = element_blank(),
panel.grid.major.y = element_line(colour = "grey50"),
plot.title = element_text(size = rel(1.5),
face = "bold", vjust = 1.5),
axis.title = element_text(face = "bold"),
legend.position = "top",
legend.title = element_blank(),
legend.key.size = unit(0.4, "cm"),
legend.key = element_rect(fill = "black"),
axis.title.y = element_text(vjust= 1.8),
axis.title.x = element_text(vjust= -0.5)
)
# # #

Bar plot of proportions vs. categories. Error bars indicate 95% confidence intervals for each observed proportion.
See the Handbook for information on these topics. The exact test of goodness-of-fit, the G-test of goodness-of-fit, and the exact test of goodness-of-fit tests are described elsewhere in this book.
How to do the test
Chi-square goodness-of-fit example
###
--------------------------------------------------------------
### Pea color example, Chi-square goodness-of-fit, pp. 50–51
### --------------------------------------------------------------
observed = c(423, 133)
expected = c(0.75, 0.25)
chisq.test(x = observed,
p = expected)
X-squared = 0.3453, df = 1, p-value = 0.5568
# # #
Power analysis
Power analysis for chi-square goodness-of-fit
###
--------------------------------------------------------------
### Power analysis, Chi-square goodness-of-fit, snapdragons, p. 51
### --------------------------------------------------------------
library(pwr)
P0 = c(0.25, 0.50, 0.25)
P1 = c(0.225, 0.55, 0.225)
effect.size = ES.w1(P0, P1)
degrees = length(P0) - 1
pwr.chisq.test(
w=effect.size,
N=NULL, # Total number of
observations
df=degrees,
power=0.80, # 1 minus Type II
probability
sig.level=0.05) # Type I
probability
N = 963.4689
# # #
![Batsto Lake, Wharton State Forest, New Jersey [banner]](images/banner.png)