Examples in Summary and Analysis of Extension Program Evaluation
SAEPER: Introduction to Parametric Tests
SAEPER:
One-way ANOVA
SAEPER: What are
Least Square Means?
Packages used in this chapter
The following commands will install these packages if they are not already installed:
if(!require(dplyr)){install.packages("dplyr")}
if(!require(FSA)){install.packages("FSA")}
if(!require(car)){install.packages("car")}
if(!require(agricolae)){install.packages("agricolae")}
if(!require(multcomp)){install.packages("multcomp")}
if(!require(DescTools)){install.packages("DescTools")}
if(!require(lsmeans)){install.packages("lsmeans")}
if(!require(multcompView)){install.packages("multcompView")}
if(!require(Rmisc)){install.packages("Rmisc")}
if(!require(ggplot2)){install.packages("ggplot2")}
if(!require(pwr)){install.packages("pwr")}
When to use it
Analysis for this example is described below in the “How to do the test” section below.
Null hypothesis
How the test works
Assumptions
Additional analyses
See the Handbook for information on these topics.
Tukey-Kramer test
The Tukey mean separation tests and others are shown below in the “How to do the test” section.
Partitioning variance
This topic is not covered here.
Example
Code for this example is not included here. An example is covered below in the “How to do the test” section.
Graphing the results
Graphing of the results is shown below in the “How to do the test” section.
Similar tests
Two-sample t–test, Two-way anova, Nested anova, Welch’s anova, and Kruskal–Wallis are presented elsewhere in this book.
A permutation test, presented in the One-way Analysis with Permutation Test chapter, can also be employed as a nonparametric alternative.
How to do the test
The lm function in the native stats package fits a linear model by least squares, and can be used for a variety of analyses such as regression, analysis of variance, and analysis of covariance. The analysis of variance is then conducted either with the Anova function in the car package for Type II or Type III sum of squares, or with the anova function in the native stats package for Type I sum of squares.
If the analysis of variance indicates a significant effect of the independent variable, multiple comparisons among the levels of this factor can be conducted using Tukey or Least Significant Difference (LSD) procedures. The problem of inflating the Type I Error Rate when making multiple comparisons is discussed in the Multiple Comparisons chapter in the Handbook. R functions which make multiple comparisons usually allow for adjusting p-values. In R, the “BH”, or “fdr”, procedure is the Benjamini–Hochberg procedure discussed in the Handbook. See ?p.adjust for more information.
One-way anova example
### --------------------------------------------------------------
### One-way anova, SAS example, pp. 155-156
### --------------------------------------------------------------
Input =("
Location Aam
Tillamook 0.0571
Tillamook 0.0813
Tillamook 0.0831
Tillamook 0.0976
Tillamook 0.0817
Tillamook 0.0859
Tillamook 0.0735
Tillamook 0.0659
Tillamook 0.0923
Tillamook 0.0836
Newport 0.0873
Newport 0.0662
Newport 0.0672
Newport 0.0819
Newport 0.0749
Newport 0.0649
Newport 0.0835
Newport 0.0725
Petersburg 0.0974
Petersburg 0.1352
Petersburg 0.0817
Petersburg 0.1016
Petersburg 0.0968
Petersburg 0.1064
Petersburg 0.1050
Magadan 0.1033
Magadan 0.0915
Magadan 0.0781
Magadan 0.0685
Magadan 0.0677
Magadan 0.0697
Magadan 0.0764
Magadan 0.0689
Tvarminne 0.0703
Tvarminne 0.1026
Tvarminne 0.0956
Tvarminne 0.0973
Tvarminne 0.1039
Tvarminne 0.1045
")
Data = read.table(textConnection(Input),header=TRUE)
Specify the order of factor levels for plots and Dunnett comparison
library(dplyr)
Data =
mutate(Data,
Location = factor(Location, levels=unique(Location)))
Produce summary statistics
library(FSA)
Summarize(Aam ~ Location,
data=Data,
digits=3)
Location n mean sd min Q1 median Q3 max
1 Tillamook 10 0.080 0.012 0.057 0.075 0.082 0.085 0.098
2 Newport 8 0.075 0.009 0.065 0.067 0.074 0.082 0.087
3 Petersburg 7 0.103 0.016 0.082 0.097 0.102 0.106 0.135
4 Magadan 8 0.078 0.013 0.068 0.069 0.073 0.081 0.103
5 Tvarminne 6 0.096 0.013 0.070 0.096 0.100 0.104 0.104
Fit the linear model and conduct ANOVA
model = lm(Aam ~ Location,
data=Data)
library(car)
Anova(model, type="II") #
Can use type="III"
### If you use type="III", you need the following line before the
analysi
### options(contrasts = c("contr.sum",
"contr.poly"))
Sum Sq Df F value Pr(>F)
Location 0.0045197 4 7.121 0.0002812 ***
Residuals 0.0053949 34
anova(model) # Produces type I sum of squares
Df Sum Sq Mean Sq F value Pr(>F)
Location 4 0.0045197 0.00112992 7.121 0.0002812 ***
Residuals 34 0.0053949 0.00015867
summary(model) # Produces r-square, overall p-value, parameter estimates
Multiple R-squared: 0.4559, Adjusted R-squared: 0.3918
F-statistic: 7.121 on 4 and 34 DF, p-value: 0.0002812
Checking assumptions of the model
hist(residuals(model),
col="darkgray")

A histogram of residuals from a linear model. The distribution of
these residuals should be approximately normal.
plot(fitted(model),
residuals(model))

A plot of residuals vs. predicted values. The residuals should be unbiased and homoscedastic. For an illustration of these properties, see this diagram by Steve Jost at DePaul University: condor.depaul.edu/sjost/it223/documents/resid-plots.gif.
### additional model checking plots
with: plot(model)
### alternative: library(FSA);
residPlot(model)
Tukey and Least Significant Difference mean separation tests (pairwise comparisons)
Tukey and other multiple comparison tests can be performed with a handful of functions. The functions TukeyHSD, HSD.test, and LSD.test are probably not appropriate for cases where there are unbalanced data or unequal variances among levels of the factor, though TukeyHSD does make an adjustment for mildly unbalanced data. It is my understanding that the multcomp and lsmeans packages are more appropriate for unbalanced data. Another alternative is the DTK package that performs mean separation tests on data with unequal sample sizes and no assumption of equal variances.
Tukey comparisons in agricolae package
library(agricolae)
(HSD.test(model, "Location")) #
outer parentheses print result
trt means M
1 Petersburg 0.1034429 a
2 Tvarminne 0.0957000 ab
3 Tillamook 0.0802000 bc
4 Magadan 0.0780125 bc
5 Newport 0.0748000 c
# Means sharing the same letter are not
significantly different
LSD comparisons in agricolae package
library(agricolae)
(LSD.test(model, "Location", # outer parentheses
print result
alpha = 0.05,
p.adj="none")) # see
?p.adjust for options
trt means M
1 Petersburg 0.1034429 a
2 Tvarminne 0.0957000 a
3 Tillamook 0.0802000 b
4 Magadan 0.0780125 b
5 Newport 0.0748000 b
# Means sharing the same letter are not
significantly different
Multiple comparisons in multcomp package
Note that “Tukey” here does not mean Tukey-adjusted comparisons. It just sets up a matrix to compare each mean to each other mean.
library(multcomp)
mc = glht(model,
mcp(Location = "Tukey"))
mcs = summary(mc, test=adjusted("single-step"))
mcs
### Adjustment options: "none",
"single-step", "Shaffer",
### "Westfall", "free",
"holm", "hochberg",
### "hommel", "bonferroni",
"BH", "BY", "fdr"
Linear Hypotheses:
Estimate Std. Error t value Pr(>|t|)
Newport - Tillamook == 0 -0.005400 0.005975 -0.904 0.89303
Petersburg - Tillamook == 0 0.023243 0.006208 3.744 0.00555 **
Magadan - Tillamook == 0 -0.002188 0.005975 -0.366 0.99596
Tvarminne - Tillamook == 0 0.015500 0.006505 2.383 0.14413
Petersburg - Newport == 0 0.028643 0.006519 4.394 < 0.001 ***
Magadan - Newport == 0 0.003213 0.006298 0.510 0.98573
Tvarminne - Newport == 0 0.020900 0.006803 3.072 0.03153 *
Magadan - Petersburg == 0 -0.025430 0.006519 -3.901 0.00376 **
Tvarminne - Petersburg == 0 -0.007743 0.007008 -1.105 0.80211
Tvarminne - Magadan == 0 0.017688 0.006803 2.600 0.09254 .
cld(mcs,
level=0.05,
decreasing=TRUE)
Tillamook Newport Petersburg Magadan Tvarminne
"bc" "c" "a" "bc" "ab"
### Means sharing a letter are
not significantly different
Multiple comparisons to a control in multcomp package
### Control is the first level of the
factor
library(multcomp)
mc = glht(model,
mcp(Location = "Dunnett"))
summary(mc, test=adjusted("single-step"))
### Adjustment options: "none",
"single-step", "Shaffer",
### "Westfall", "free",
"holm", "hochberg",
### "hommel", "bonferroni",
"BH", "BY", "fdr"
Linear Hypotheses:
Estimate Std. Error t value Pr(>|t|)
Newport - Tillamook == 0 -0.005400 0.005975 -0.904 0.79587
Petersburg - Tillamook == 0 0.023243 0.006208 3.744 0.00252 **
Magadan - Tillamook == 0 -0.002188 0.005975 -0.366 0.98989
Tvarminne - Tillamook == 0 0.015500 0.006505 2.383 0.07794 .
Multiple comparisons to a control with Dunnett Test
### The control group can be specified
with the control option,
### or will be the first level of the factor
library(DescTools)
DunnettTest(Aam ~ Location,
data = Data)
Dunnett's test for comparing several treatments with a control :
95% family-wise confidence level
diff lwr.ci upr.ci pval
Newport-Tillamook -0.00540000 -0.020830113 0.01003011 0.7958
Petersburg-Tillamook 0.02324286 0.007212127 0.03927359 0.0026 **
Magadan-Tillamook -0.00218750 -0.017617613 0.01324261 0.9899
Tvarminne-Tillamook 0.01550000 -0.001298180 0.03229818 0.0778 .
Multiple comparisons with least square means
Least square means can be calculated for each group. Here a Tukey adjustment is applied for multiple comparisons among group least square means. The multiple comparisons can be displayed as a compact letter display.
library(lsmeans)
library(multcompView)
leastsquare = lsmeans(model,
pairwise ~ Location,
adjust = "tukey")
$contrasts
contrast estimate SE df t.ratio p.value
Tillamook - Newport 0.005400000 0.005975080 34 0.904 0.8935
Tillamook - Petersburg -0.023242857 0.006207660 34 -3.744 0.0057
Tillamook - Magadan 0.002187500 0.005975080 34 0.366 0.9960
Tillamook - Tvarminne -0.015500000 0.006504843 34 -2.383 0.1447
Newport - Petersburg -0.028642857 0.006519347 34 -4.394 0.0009
Newport - Magadan -0.003212500 0.006298288 34 -0.510 0.9858
Newport - Tvarminne -0.020900000 0.006802928 34 -3.072 0.0317
Petersburg - Magadan 0.025430357 0.006519347 34 3.901 0.0037
Petersburg - Tvarminne 0.007742857 0.007008087 34 1.105 0.8028
Magadan - Tvarminne -0.017687500 0.006802928 34 -2.600 0.0929
P value adjustment: tukey method for comparing a family of 5 estimates
cld(leastsquare,
alpha = 0.05,
Letters = letters,
adjust="tukey")
Location lsmean SE df lower.CL upper.CL .group
Newport 0.0748000 0.004453562 34 0.06268565 0.08691435 a
Magadan 0.0780125 0.004453562 34 0.06589815 0.09012685 ab
Tillamook 0.0802000 0.003983387 34 0.06936459 0.09103541 ab
Tvarminne 0.0957000 0.005142530 34 0.08171155 0.10968845 bc
Petersburg 0.1034429 0.004761058 34 0.09049207 0.11639365 c
Confidence level used: 0.95
Conf-level adjustment: sidak method for 5 estimates
P value adjustment: tukey method for comparing a family of 5 estimates
significance level used: alpha = 0.05
Graphing the results
Simple box plots of values across groups
boxplot(Aam ~ Location,
data = Data,
ylab="aam / height",
xlab="Location")
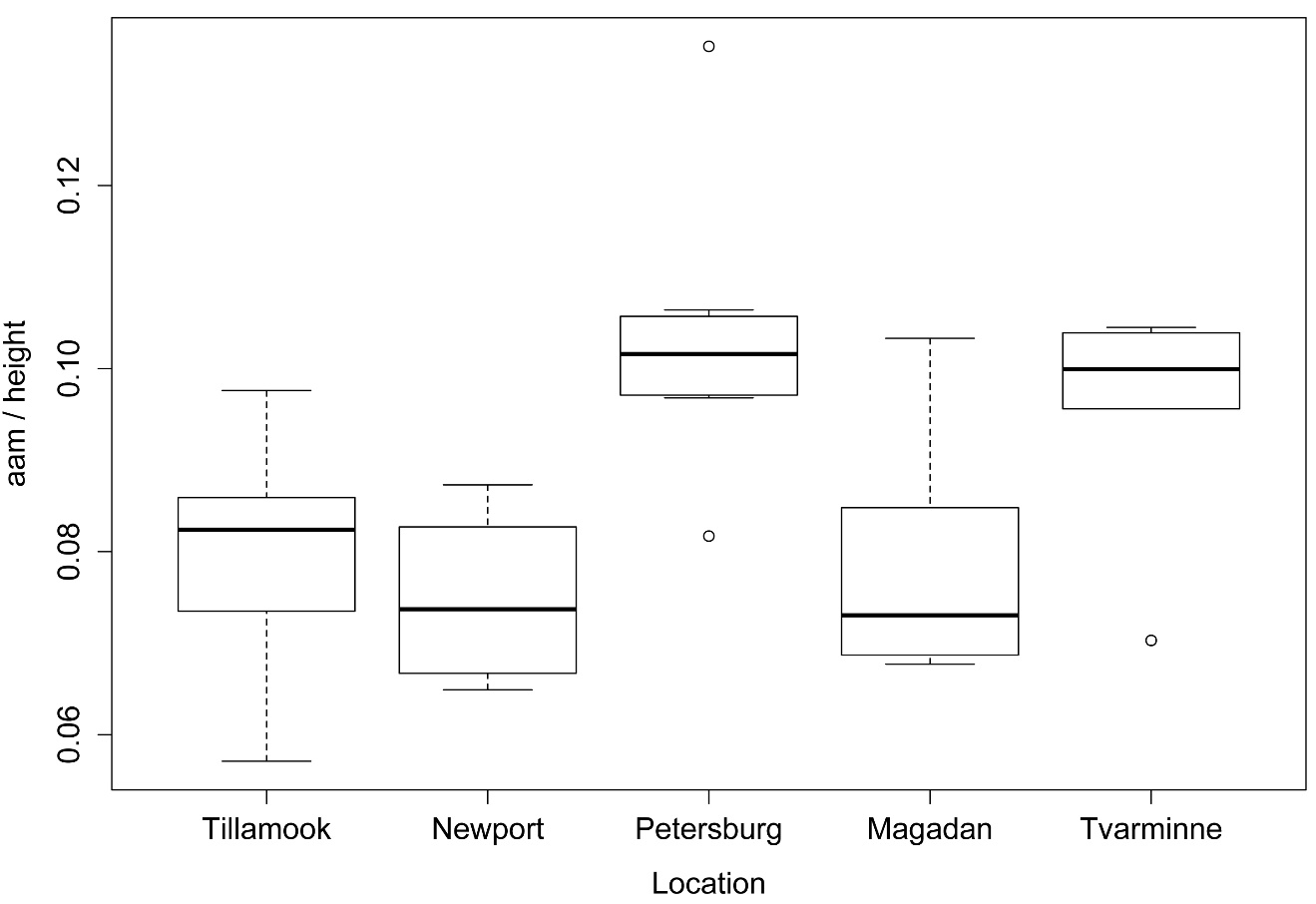
Box plots of values for each level of the independent variable for a one-way analysis of variance (ANOVA).
Simple bar plot of means across groups
### Summarize the data frame (Data)
into a table
library(Rmisc)
Data2 = summarySE(data=Data,
"Aam",
groupvars="Location",
conf.interval = 0.95)
Tabla = as.table(Data2$Aam)
rownames(Tabla) = Data2$Location
Tabla
Tillamook Newport Petersburg Magadan Tvarminne
0.0802000 0.0748000 0.1034429 0.0780125 0.0957000
barplot(Tabla,
ylab="aam / height",
xlab="Location")

Bar plot of means for each level of the independent variable for a one-way analysis of variance (ANOVA).
Bar plot of means with error bars across groups
library(ggplot2)
offset.v = -3 # offsets for mean letters
offset.h
= 0.5
ggplot(Data2,
aes(x = Location, y = Aam,
ymax=0.12, ymin=0.0)) +
geom_bar(stat="identity", fill="gray50",
colour = "black", width = 0.7) +
geom_errorbar(aes(ymax=Aam+se, ymin=Aam-se),
width=0.0, size=0.5, color="black") +
geom_text(aes(label=c("bc","c","a","bc","ab"),
hjust=offset.h, vjust=offset.v)) +
labs(x = "Sample location",
y = "aam / height") +
## ggtitle("Main title") +
theme_bw() +
theme(panel.grid.major.x = element_blank(),
panel.grid.major.y = element_line(colour = "grey80"),
plot.title = element_text(size = rel(1.5),
face = "bold", vjust = 1.5),
axis.title = element_text(face = "bold"),
axis.title.y = element_text(vjust= 1.8),
axis.title.x = element_text(vjust= -0.5),
panel.border = element_rect(colour="black")
)

Bar plot of means for each level of the independent variable of a one-way analysis of variance (ANOVA). Error indicates standard error of the mean. Bars sharing the same letter are not significantly different according to Tukey’s HSD test.
Welch’s anova
Bartlett’s test and Levene’s test can be used to check the homoscedasticity of groups from a one-way anova. A significant result for these tests (p < 0.05) suggests that groups are heteroscedastic. One approach with heteroscedastic data in a one way anova is to use the Welch correction with the oneway.test function in the native stats package. A more versatile approach is to use the white.adjust=TRUE option in the Anova function from the car package.
### Bartlett test for homogeneity of variance
bartlett.test(Aam ~ Location,
data = Data)
Bartlett test of homogeneity of variances
Bartlett's K-squared = 2.4341, df = 4, p-value = 0.6565
### Levene test for homogeneity of
variance
library(car)
leveneTest(Aam ~ Location,
data = Data)
Levene's Test for Homogeneity of Variance (center = median)
Df F value Pr(>F)
group 4 0.12 0.9744
34
### Welch’s anova for unequal variances
oneway.test(Aam ~ Location,
data=Data,
var.equal=FALSE)
One-way analysis of means (not assuming equal variances)
F = 5.6645, num df = 4.000, denom df = 15.695, p-value = 0.00508
### White-adjusted anova for
heteroscedasticity
model = lm(Aam ~ Location,
data=Data)
library(car)
Anova(model, Type="II",
white.adjust=TRUE)
Df F Pr(>F)
Location 4 5.4617 0.001659 **
Residuals 34
# # #
Power analysis
Power analysis for one-way anova
###
--------------------------------------------------------------
### Power analysis for anova, pp. 157
### --------------------------------------------------------------
library(pwr)
groups = 5
means = c(10, 10, 15, 15, 15)
sd = 12
grand.mean = mean(means)
Cohen.f = sqrt( sum( (1/groups) * (means-grand.mean)^2) ) /sd
pwr.anova.test(k = groups,
n = NULL,
f = Cohen.f,
sig.level = 0.05,
power = 0.80)
Balanced one-way analysis of variance power calculation
n = 58.24599
NOTE: n is number in each group
# # #
![Batsto Lake, Wharton State Forest, New Jersey [banner]](images/banner.png)