A two-way ordinal analysis of variance can address an
experimental design with two independent variables, each of which is a factor
variable. The main effect of each independent variable can be tested, as well
as the effect of the interaction of the two factors.
The example here looks at ratings for three instructors across four different questions. The analysis will attempt to answer the questions: a) Is there a significant difference in scores for different instructors? b) Is there a significant difference in scores for different questions? c) Is there a significant interaction effect of Instructor and Question?
Main effects and interaction effects are explained further
in the Factorial ANOVA: Main Effects, Interaction Effects, and Interaction Plots
chapter.
The clm function can specify more complex models with
multiple independent variables of different types, but this book will not explore
more complex examples.
Post-hoc analysis to determine which groups are different can
be conducted on each significant main effect and on the interaction effect if
it is significant.
Appropriate data
• Two-way data
• Dependent variable is ordered factor
• Independent variables are factors with at least two levels or groups each
• Observations between groups are not paired or repeated measures
Interpretation
A significant main effect can be interpreted as, “There was a
significant difference among groups.” Or, “There was a significant effect of
Independent Variable.”
A significant interaction effect can be interpreted as, “There
was a significant interaction effect between Independent Variable A and
Independent Variable B.”
A significant post-hoc analysis indicates, “There was a
significant difference between Group A and Group B”, and so on.
Packages used in this chapter
The packages used in this chapter include:
• psych
• FSA
• lattice
• ggplot2
• ordinal
• car
• RVAideMemoire
• rcompanion
• multcomp
• emmeans
The following commands will install these packages if they are not already installed:
if(!require(psych)){install.packages("psych")}
if(!require(FSA)){install.packages("FSA")}
if(!require(lattice)){install.packages("lattice")}
if(!require(ggplot2)){install.packages("ggplot2")}
if(!require(ordinal)){install.packages("ordinal")}
if(!require(car)){install.packages("car")}
if(!require(RVAideMemoire)){install.packages("RVAideMemoire")}
if(!require(rcompanion)){install.packages("rcompanion")}
if(!require(multcomp)){install.packages("multcomp")}
if(!require(emmeans)){install.packages("emmeans")}
Two-way ordinal regression example
Data = read.table(header=TRUE, stringsAsFactors=TRUE, text="
Instructor Question Likert
Fuu Informative 8
Fuu Informative 9
Fuu Informative 9
Fuu Informative 8
Fuu Delivery 10
Fuu Delivery 9
Fuu Delivery 8
Fuu Delivery 8
Fuu VisualAides 7
Fuu VisualAides 7
Fuu VisualAides 6
Fuu VisualAides 7
Fuu AnswerQuest 8
Fuu AnswerQuest 9
Fuu AnswerQuest 9
Fuu AnswerQuest 8
Jin Informative 7
Jin Informative 8
Jin Informative 6
Jin Informative 5
Jin Delivery 8
Jin Delivery 8
Jin Delivery 9
Jin Delivery 6
Jin VisualAides 5
Jin VisualAides 6
Jin VisualAides 7
Jin VisualAides 7
Jin AnswerQuest 8
Jin AnswerQuest 7
Jin AnswerQuest 6
Jin AnswerQuest 6
Mugen Informative 5
Mugen Informative 4
Mugen Informative 3
Mugen Informative 4
Mugen Delivery 8
Mugen Delivery 9
Mugen Delivery 8
Mugen Delivery 7
Mugen VisualAides 5
Mugen VisualAides 4
Mugen VisualAides 4
Mugen VisualAides 5
Mugen AnswerQuest 6
Mugen AnswerQuest 7
Mugen AnswerQuest 6
Mugen AnswerQuest 7
")
### Order levels of the factor; otherwise R will alphabetize them
Data$Instructor = factor(Data$Instructor,
levels=unique(Data$Instructor))
### Create a new variable which is the likert scores as an ordered factor
Data$Likert.f = factor(Data$Likert,
ordered = TRUE)
### Check the data frame
library(psych)
headTail(Data)
str(Data)
summary(Data)
Summarize data treating Likert scores as factors
xtabs( ~ Instructor + Likert.f,
data = Data)
Likert.f
Instructor 3 4 5 6 7 8 9 10
Fuu 0 0 0 1 3 6 5 1
Jin 0 0 2 5 4 4 1 0
Mugen 1 4 3 2 3 2 1 0
xtabs( ~ Question + Likert.f,
data = Data)
Likert.f
Question 3 4 5 6 7 8 9 10
AnswerQuest 0 0 0 4 3 3 2 0
Delivery 0 0 0 1 1 6 3 1
Informative 1 2 2 1 1 3 2 0
VisualAides 0 2 3 2 5 0 0 0
xtabs( ~ Instructor + Likert.f + Question,
data = Data)
, , Question = AnswerQuest
Likert.f
Instructor 3 4 5 6 7 8 9 10
Fuu 0 0 0 0 0 2 2 0
Jin 0 0 0 2 1 1 0 0
Mugen 0 0 0 2 2 0 0 0
, , Question = Delivery
Likert.f
Instructor 3 4 5 6 7 8 9 10
Fuu 0 0 0 0 0 2 1 1
Jin 0 0 0 1 0 2 1 0
Mugen 0 0 0 0 1 2 1 0
, , Question = Informative
Likert.f
Instructor 3 4 5 6 7 8 9 10
Fuu 0 0 0 0 0 2 2 0
Jin 0 0 1 1 1 1 0 0
Mugen 1 2 1 0 0 0 0 0
, , Question = VisualAides
Likert.f
Instructor 3 4 5 6 7 8 9 10
Fuu 0 0 0 1 3 0 0 0
Jin 0 0 1 1 2 0 0 0
Mugen 0 2 2 0 0 0 0 0
Bar plots by group
library(lattice)
histogram(~ Likert.f | Instructor,
data=Data,
layout=c(1,3) # columns and rows of
individual plots
)

library(lattice)
histogram(~ Likert.f | Question,
data=Data,
layout=c(1,4) # columns and rows of
individual plots
)

library(lattice)
histogram(~ Likert.f | Instructor + Question,
data=Data,
layout=c(3,4) # columns and rows of
individual plots
)

Summarize data treating Likert scores as numeric
library(FSA)
Summarize(Likert ~ Instructor,
data=Data,
digits=3)
Instructor n mean sd min Q1 median Q3 max
percZero
1 Fuu 16 8.125 1.025 6 7.75 8.0 9 10 0
2 Jin 16 6.812 1.167 5 6.00 7.0 8 9 0
3 Mugen 16 5.750 1.770 3 4.00 5.5 7 9 0
library(FSA)
Summarize(Likert ~ Question,
data=Data,
digits=3)
Question n mean sd min Q1 median Q3 max percZero
1 AnswerQuest 12 7.250 1.138 6 6.00 7.0 8 9 0
2 Delivery 12 8.167 1.030 6 8.00 8.0 9 10 0
3 Informative 12 6.333 2.103 3 4.75 6.5 8 9 0
4 VisualAides 12 5.833 1.193 4 5.00 6.0 7 7 0
library(FSA)
Summarize(Likert ~ Instructor + Question,
data=Data,
digits=3)
Instructor Question n mean sd min Q1 median Q3 max
percZero
1 Fuu AnswerQuest 4 8.50 0.577 8 8.00 8.5 9.00 9 0
2 Jin AnswerQuest 4 6.75 0.957 6 6.00 6.5 7.25 8 0
3 Mugen AnswerQuest 4 6.50 0.577 6 6.00 6.5 7.00 7 0
4 Fuu Delivery 4 8.75 0.957 8 8.00 8.5 9.25 10 0
5 Jin Delivery 4 7.75 1.258 6 7.50 8.0 8.25 9 0
6 Mugen Delivery 4 8.00 0.816 7 7.75 8.0 8.25 9 0
7 Fuu Informative 4 8.50 0.577 8 8.00 8.5 9.00 9 0
8 Jin Informative 4 6.50 1.291 5 5.75 6.5 7.25 8 0
9 Mugen Informative 4 4.00 0.816 3 3.75 4.0 4.25 5 0
10 Fuu VisualAides 4 6.75 0.500 6 6.75 7.0 7.00 7 0
11 Jin VisualAides 4 6.25 0.957 5 5.75 6.5 7.00 7 0
12 Mugen VisualAides 4 4.50 0.577 4 4.00 4.5 5.00 5 0
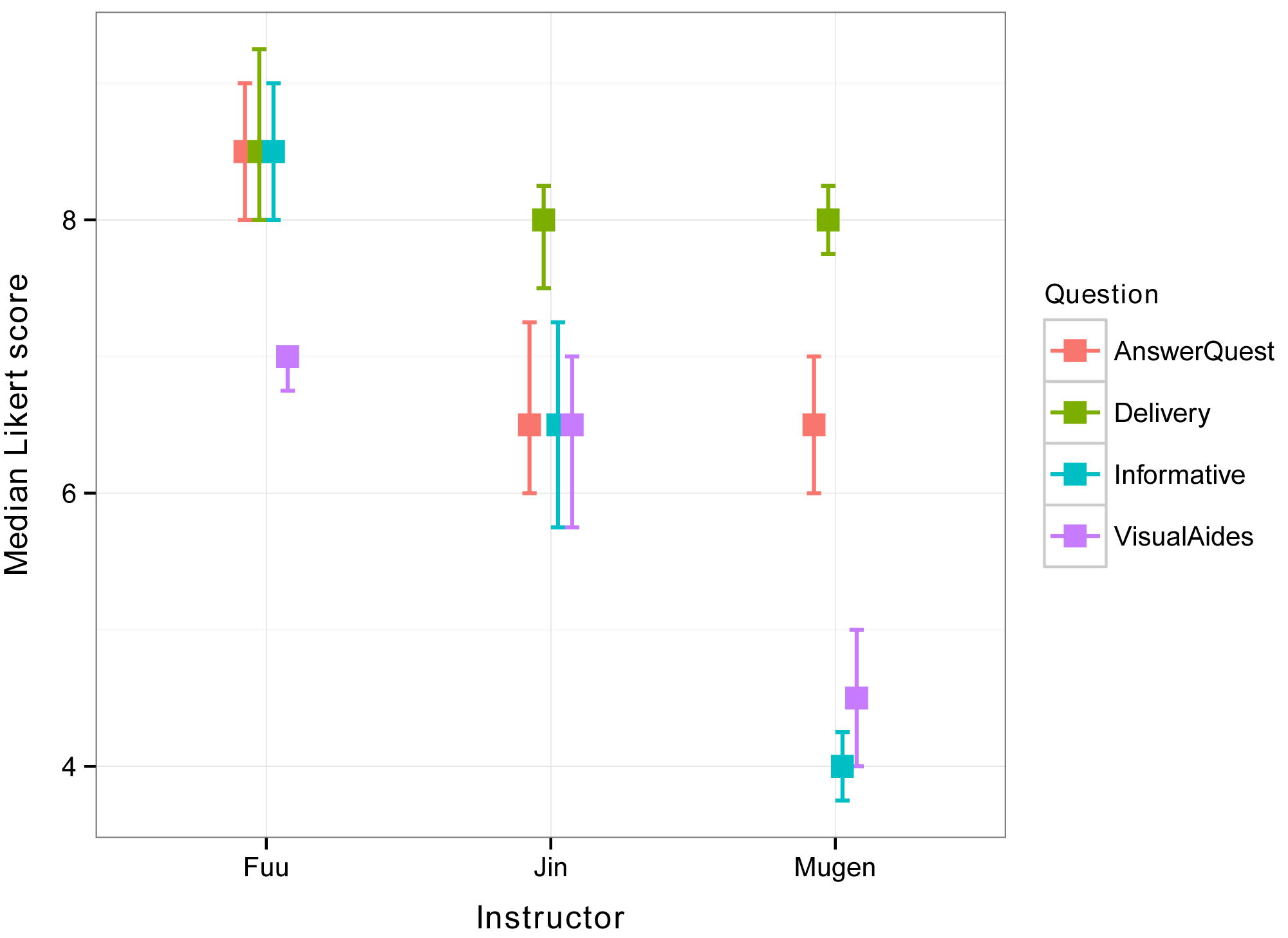
Produce interaction plot with medians and quantiles
The following plot shows medians for the interaction of Instructor and Question, with error bars showing the 1st quartile and 3rd quartile for each median.
library(FSA)
Sum = Summarize(Likert ~ Instructor + Question,
data=Data,
digits=3)
Sum
library(ggplot2)
pd = position_dodge(.2)
ggplot(Sum, aes(x=Instructor,
y=median,
color=Question)) +
geom_errorbar(aes(ymin=Q1,
ymax=Q3),
width=.2, size=0.7, position=pd) +
geom_point(shape=15, size=4, position=pd) +
theme_bw() +
theme(axis.title = element_text(face = "bold"))+
ylab("Median Likert score")
Two-way ordinal regression
In the model notation in the clm function, here, Likert.f
is the dependent variable and Instructor and Question are the
independent variables. The term Instructor:Question adds the
interaction effect of these two independent variables to the model. The data=
option indicates the data frame that contains the variables. For the meaning
of other options, see ?clm.
Here the threshold = "symmetric" option is
used in order to avoid errors. This option does not need to be used routinely.
Define model and analyses of deviance
library(ordinal)
model = clm(Likert.f ~ Instructor + Question + Instructor:Question,
data = Data,
threshold="symmetric")
Analysis of deviance
anova(model, type="II")
Type II Analysis of Deviance Table with Wald chi-square tests
Df Chisq Pr(>Chisq)
Instructor 2 26.748 1.555e-06 ***
Question 3 24.596 1.876e-05 ***
Instructor:Question 6 18.194 0.005766 **
Alternate analysis of deviance analysis
library(car)
library(RVAideMemoire)
Anova.clm(model, type = "II")
Analysis of Deviance Table (Type II tests)
LR Chisq Df Pr(>Chisq)
Instructor 32.157 2 1.040e-07 ***
Question 28.248 3 3.221e-06 ***
Instructor:Question 24.326 6 0.0004548 ***
p-value for model and pseudo R-squared
The p-value for the model and a pseudo R-squared value can be determined with the nagelkerke function. It is not clear to me which pseudo R-squared value would be most appropriate for an ordinal regression model.
library(rcompanion)
nagelkerke(model)
$Pseudo.R.squared.for.model.vs.null
Pseudo.R.squared
McFadden 0.400602
Cox and Snell (ML) 0.775956
Nagelkerke (Cragg and Uhler) 0.794950
$Likelihood.ratio.test
Df.diff LogLik.diff Chisq p.value
-11 -35.902 71.804 5.5398e-11
Post-hoc tests
For clm model objects, the emmean, SE, LCL,
and UCL values should be ignored, unless specific options in emmeans
are selected.
Because the interaction term in the model was significant, the group separation for the interaction effect is explored.
library(emmeans)
marginal = emmeans(model,
~ Instructor + Question)
pairs(marginal,
adjust="tukey")
library(multcomp)
cld(marginal, Letters=letters)
Instructor Question emmean SE df asymp.LCL asymp.UCL .group
Mugen Informative -6.663 1.419 Inf -9.444 -3.88 a
Mugen VisualAides -5.263 1.279 Inf -7.770 -2.76 ab
Jin VisualAides -0.435 0.944 Inf -2.284 1.41 bc
Jin Informative 0.000 1.055 Inf -2.067 2.07 bcd
Mugen AnswerQuest 0.000 0.848 Inf -1.663 1.66 c
Jin AnswerQuest 0.435 0.944 Inf -1.415 2.28 cde
Fuu VisualAides 0.553 0.846 Inf -1.107 2.21 cd
Jin Delivery 3.490 1.319 Inf 0.904 6.08 cde
Mugen Delivery 3.713 1.225 Inf 1.311 6.11 cde
Fuu AnswerQuest 5.263 1.279 Inf 2.756 7.77 de
Fuu Informative 5.263 1.279 Inf 2.756 7.77 de
Fuu Delivery 5.783 1.378 Inf 3.082 8.48 e
Confidence level used: 0.95
P value adjustment: tukey method for comparing a family of 12 estimates
significance level used: alpha = 0.05
NOTE: Compact letter displays can be misleading
because they show NON-findings rather than findings.
Consider using 'pairs()', 'pwpp()', or 'pwpm()' instead.
### Groups sharing a letter in .group are not
significantly different
### Remember to ignore “emmean”, “SE”, “LCL”, and
“UCL” with CLM
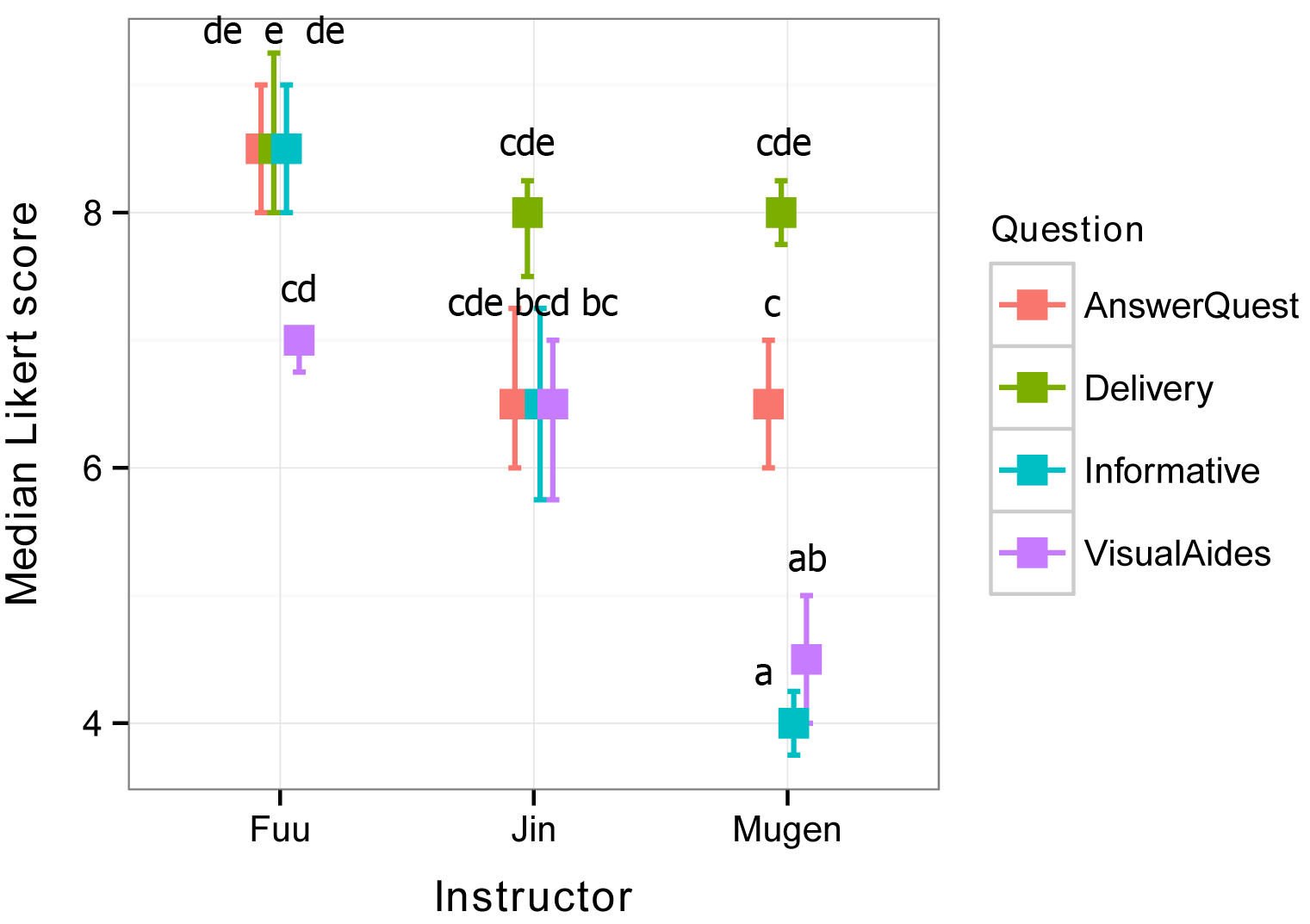
Interaction plot with group separation letters
Group separation letters can be added manually to an
interaction plot.
Groups sharing a letter are not significantly different. In
this case, because so many groups share a letter, it is difficult to interpret
this plot. One approach would be to look at differences among instructors for
each question. Looking at AnswerQuest, Fuu’s scores are not
statistically different than Jin’s (because they share the letters d and
e), but Fuu’s scores are different than Mugen’s (because they share no
letters). So, we can conclude for this question, that Fuu’s scores are
significantly greater than Mugen’s. And so on.
Check model assumptions
nominal_test(model)
Tests of nominal effects
formula: Likert.f ~ Instructor + Question + Instructor:Question
Df logLik AIC LRT Pr(>Chi)
<none> -53.718 137.44
Instructor 6 -49.812 141.62 7.8121 0.2522
Question 9 -50.182 148.37 7.0711 0.6297
Instructor:Question
### No violation in assumptions.
scale_test(model)
Tests of scale effects
formula: Likert.f ~ Instructor + Question + Instructor:Question
Df logLik AIC LRT Pr(>Chi)
<none> -53.718 137.44
Instructor 2 -51.669 137.34 4.0985 0.12883
Question 3 -50.534 137.07 6.3669 0.09506 .
Instructor:Question
Warning message:
(-1) Model failed to converge with max|grad| = 1.70325e-06 (tol = 1e-06)
In addition: maximum number of consecutive Newton modifications reached
### This test failed, but the results suggest no
violation of assumptions.
![Cohansey River, Fairfield, New Jersey [banner]](images/banner.jpg)